|
In mathematics, the Stiefel manifoldVk(Rn){displaystyle V_{k}(mathbb {R} ^{n})} is the set of all orthonormalk-frames in Rn.{displaystyle mathbb {R} ^{n}.} That is, it is the set of ordered k-tuples of orthonormal vectors in Rn.{displaystyle mathbb {R} ^{n}.} It is named after Swiss mathematician Eduard Stiefel. Likewise one can define the complex Stiefel manifold Vk(Cn){displaystyle V_{k}(mathbb {C} ^{n})} of orthonormal k-frames in Cn{displaystyle mathbb {C} ^{n}} and the quaternionic Stiefel manifold Vk(Hn){displaystyle V_{k}(mathbb {H} ^{n})} of orthonormal k-frames in Hn{displaystyle mathbb {H} ^{n}} More generally, the construction applies to any real, complex, or quaternionic inner product space.
In some contexts, a non-compact Stiefel manifold is defined as the set of all linearly independentk-frames in Rn,Cn,{displaystyle mathbb {R} ^{n},mathbb {C} ^{n},} or Hn;{displaystyle mathbb {H} ^{n};} this is homotopy equivalent, as the compact Stiefel manifold is a deformation retract of the non-compact one, by Gram–Schmidt. Statements about the non-compact form correspond to those for the compact form, replacing the orthogonal group (or unitary or symplectic group) with the general linear group.
Topology[edit]
Let F{displaystyle mathbb {F} } stand for R,C,{displaystyle mathbb {R} ,mathbb {C} ,} or H.{displaystyle mathbb {H} .} The Stiefel manifold Vk(Fn){displaystyle V_{k}(mathbb {F} ^{n})} can be thought of as a set of n × kmatrices by writing a k-frame as a matrix of kcolumn vectors in Fn.{displaystyle mathbb {F} ^{n}.} The orthonormality condition is expressed by A*A = Ik{displaystyle I_{k}} where A* denotes the conjugate transpose of A and Ik{displaystyle I_{k}} denotes the k × kidentity matrix. We then have
The topology on Vk(Fn){displaystyle V_{k}(mathbb {F} ^{n})} is the subspace topology inherited from Fn×k.{displaystyle mathbb {F} ^{ntimes k}.} With this topology Vk(Fn){displaystyle V_{k}(mathbb {F} ^{n})} is a compactmanifold whose dimension is given by
As a homogeneous space[edit]
Each of the Stiefel manifolds Vk(Fn){displaystyle V_{k}(mathbb {F} ^{n})} can be viewed as a homogeneous space for the action of a classical group in a natural manner.
Every orthogonal transformation of a k-frame in Rn{displaystyle mathbb {R} ^{n}} results in another k-frame, and any two k-frames are related by some orthogonal transformation. In other words, the orthogonal group O(n) acts transitively on Vk(Rn).{displaystyle V_{k}(mathbb {R} ^{n}).} The stabilizer subgroup of a given frame is the subgroup isomorphic to O(n−k) which acts nontrivially on the orthogonal complement of the space spanned by that frame.
Likewise the unitary group U(n) acts transitively on Vk(Cn){displaystyle V_{k}(mathbb {C} ^{n})} with stabilizer subgroup U(n−k) and the symplectic group Sp(n) acts transitively on Vk(Hn){displaystyle V_{k}(mathbb {H} ^{n})} with stabilizer subgroup Sp(n−k).
In each case Vk(Fn){displaystyle V_{k}(mathbb {F} ^{n})} can be viewed as a homogeneous space:
When k = n, the corresponding action is free so that the Stiefel manifold Vn(Fn){displaystyle V_{n}(mathbb {F} ^{n})} is a principal homogeneous space for the corresponding classical group.
When k is strictly less than n then the special orthogonal group SO(n) also acts transitively on Vk(Rn){displaystyle V_{k}(mathbb {R} ^{n})} with stabilizer subgroup isomorphic to SO(n−k) so that
The same holds for the action of the special unitary group on Vk(Cn){displaystyle V_{k}(mathbb {C} ^{n})}
Thus for k = n − 1, the Stiefel manifold is a principal homogeneous space for the corresponding special classical group.
Uniform measure[edit]
The Stiefel manifold can be equipped with a uniform measure, i.e. a Borel measure that is invariant under the action of the groups noted above. For example, V1(R2){displaystyle V_{1}(mathbb {R} ^{2})} which is isomorphic to the unit circle in the Euclidean plane, has as its uniform measure the obvious uniform measure (arc length) on the circle. It is straightforward to sample this measure on Vk(Fn){displaystyle V_{k}(mathbb {F} ^{n})} using Gaussian random matrices: if A∈Fn×k{displaystyle Ain mathbb {F} ^{ntimes k}} is a random matrix with independent entries identically distributed according to the standard normal distribution on F{displaystyle mathbb {F} } and A = QR is the QR factorization of A, then the matrices, Q∈Fn×k,R∈Fk×k{displaystyle Qin mathbb {F} ^{ntimes k},Rin mathbb {F} ^{ktimes k}} are independent random variables and Q is distributed according to the uniform measure on Vk(Fn).{displaystyle V_{k}(mathbb {F} ^{n}).} This result is a consequence of the Bartlett decomposition theorem.[1]
Special cases[edit]
A 1-frame in Fn{displaystyle mathbb {F} ^{n}} is nothing but a unit vector, so the Stiefel manifold V1(Fn){displaystyle V_{1}(mathbb {F} ^{n})} is just the unit sphere in Fn.{displaystyle mathbb {F} ^{n}.} Therefore:
Given a 2-frame in Rn,{displaystyle mathbb {R} ^{n},} let the first vector define a point in Sn−1 and the second a unit tangent vector to the sphere at that point. In this way, the Stiefel manifold V2(Rn){displaystyle V_{2}(mathbb {R} ^{n})} may be identified with the unit tangent bundleto Sn−1.
When k = n or n−1 we saw in the previous section that Vk(Fn){displaystyle V_{k}(mathbb {F} ^{n})} is a principal homogeneous space, and therefore diffeomorphic to the corresponding classical group:
Functoriality[edit]
Given an orthogonal inclusion between vector spaces X↪Y,{displaystyle Xhookrightarrow Y,} the image of a set of k orthonormal vectors is orthonormal, so there is an induced closed inclusion of Stiefel manifolds, Vk(X)↪Vk(Y),{displaystyle V_{k}(X)hookrightarrow V_{k}(Y),} and this is functorial. More subtly, given an n-dimensional vector space X, the dual basis construction gives a bijection between bases for X and bases for the dual space X∗,{displaystyle X^{*},} which is continuous, and thus yields a homeomorphism of top Stiefel manifolds Vn(X)→∼Vn(X∗).{displaystyle V_{n}(X){stackrel {sim }{to }}V_{n}(X^{*}).} This is also functorial for isomorphisms of vector spaces.
As a principal bundle[edit]
There is a natural projection
from the Stiefel manifold Vk(Fn){displaystyle V_{k}(mathbb {F} ^{n})} to the Grassmannian of k-planes in Fn{displaystyle mathbb {F} ^{n}} which sends a k-frame to the subspace spanned by that frame. The fiber over a given point P in Gk(Fn){displaystyle G_{k}(mathbb {F} ^{n})} is the set of all orthonormal k-frames contained in the space P.
This projection has the structure of a principal G-bundle where G is the associated classical group of degree k. Take the real case for concreteness. There is a natural right action of O(k) on Vk(Rn){displaystyle V_{k}(mathbb {R} ^{n})} which rotates a k-frame in the space it spans. This action is free but not transitive. The orbits of this action are precisely the orthonormal k-frames spanning a given k-dimensional subspace; that is, they are the fibers of the map p. Similar arguments hold in the complex and quaternionic cases.
We then have a sequence of principal bundles:
The vector bundlesassociated to these principal bundles via the natural action of G on Fk{displaystyle mathbb {F} ^{k}} are just the tautological bundles over the Grassmannians. In other words, the Stiefel manifold Vk(Fn){displaystyle V_{k}(mathbb {F} ^{n})} is the orthogonal, unitary, or symplectic frame bundle associated to the tautological bundle on a Grassmannian.
When one passes to the n→∞{displaystyle nto infty } limit, these bundles become the universal bundles for the classical groups.
Homotopy[edit]
The Stiefel manifolds fit into a family of fibrations:
thus the first non-trivial homotopy group of the space Vk(Rn){displaystyle V_{k}(mathbb {R} ^{n})} is in dimension n − k. Moreover,
This result is used in the obstruction-theoretic definition of Stiefel–Whitney classes.
See also[edit]References[edit]
Retrieved from 'https://en.wikipedia.org/w/index.php?title=Stiefel_manifold&oldid=896581762'
Abstract
We consider the problem of sampling from posterior distributions for Bayesian models where some parameters are restricted to be orthogonal matrices. Such matrices are sometimes used in neural networks models for reasons of regularization and stabilization of training procedures, and also can parameterize matrices of bounded rank, positive-definite matrices and others. In Byrne & Girolami (2013) authors have already considered sampling from distributions over manifolds using exact geodesic flows in a scheme similar to Hamiltonian Monte Carlo (HMC). We propose new sampling scheme for a set of orthogonal matrices that is based on the same approach, uses ideas of Riemannian optimization and does not require exact computation of geodesic flows. The method is theoretically justified by proof of symplecticity for the proposed iteration. In experiments we show that the new scheme is comparable or faster in time per iteration and more sample-efficient comparing to conventional HMC with explicit orthogonal parameterization and Geodesic Monte-Carlo. We also provide promising results of Bayesian ensembling for orthogonal neural networks and low-rank matrix factorization.
oddsidemargin has been altered.
marginparsep has been altered. topmargin has been altered. marginparwidth has been altered. marginparpush has been altered. paperheight has been altered. The page layout violates the ICML style.Please do not change the page layout, or include packages like geometry,savetrees, or fullpage, which change it for you.We’re not able to reliably undo arbitrary changes to the style. Please removethe offending package(s), or layout-changing commands and try again.
Hamiltonian Monte-Carlo for Orthogonal Matrices
Viktor Yanush0 0 Dmitry Kropotov0 0  ††footnotetext: 1AUTHORERR: Missing icmlaffiliation.2AUTHORERR: Missing icmlaffiliation..Correspondence to: Viktor Yanush <[email protected]>.@xsect
In the paper we consider the problem of sampling from posterior distributions for Bayesian models where some parameters are restricted to be orthogonal matrices. Examples of such models include orthogonal recurrent neural network (Arjovsky et al., 2016), where orthogonal hidden-to-hidden matrix is used to facilitate problems with exploding or vanishing gradients, orthogonal feed-forward neural network (Huang et al., 2017), where orthogonal matrices in fully-connected or convolutional layers give additional regularization and stabilize neural network training. Besides, orthogonal matrices can be used in parameterizations for matrices of bounded rank, positive-definite matrices or tensor decompositions.
It is known that a set of orthogonal matrices forms a Riemannian manifold (Tagare, 2011). From optimization point of view, using the properties of such manifolds can accelerate optimization procedures in many cases. Examples include optimization w.r.t. low-rank matrices (Vandereycken, 2013) and tensors in tensor train format (Steinlechner, 2015), K-FAC approach for training neural networks of different architectures (Martens & Grosse, 2015; Martens et al., 2018) and many others. Hence, it is desirable to consider the Riemannian manifold structure of orthogonal matrices within sampling procedures.
One of the major approaches for sampling in Bayesian models is Hamiltonian Monte Carlo (HMC) (Neal et al., 2011). It can be applied in stochastic setting (Chen et al., 2014) and can consider Riemannian geometry (Ma et al., 2015). However, application of these methods for sampling w.r.t. orthogonal matrices requires explicit parameterization of the corresponding manifold. In case of quadratic matrices one possible choice of unambiguous parameterization is given in (Lucas, 2011). However, it requires computing of matrix exponential which is impractical for large matrices. Another option is to consider parameterization Q=X(XTX)−1/2, where X is arbitrary [possibly rectangular] matrix. This parameterization is non-unique and in practice, as we show in this paper, could lead to slow distribution exploration.
In this paper we propose a new sampling method for Bayesian models with orthogonal matrices. Here we extend HMC approach using ideas from Riemannian optimization for orthogonal matrices from (Tagare, 2011). In general outline the main idea of the proposed method is the following. Basic Riemannian HMC approach (Girolami & Calderhead, 2011) supposes introduction of auxiliary momentum variables that are restricted to lie in tangent space for the current distribution parameters. Given some transformation of these parameters we consider vector transport transformation that maps momentum variables to tangent space for the new parameter values.Our method is an extension of previously known Geodesic Monte-Carlo (Geodesic HMC, gHMC) (Byrne & Girolami, 2013). The major difference is that step along the geodesic is replaced with retraction step which is much cheaper in terms of computation.We prove that the proposed transformation of parameters and momentum variables is symplectic – a sufficient condition for correctness of HMC sampling procedure.
We show in experiments that the proposed sampling method is way more sample-efficient than standard HMC with parameterization Q=X(XTX)−1/2. This is true both for non-stochastic and stochastic regimes. We also consider Bayesian orthogonal neural networks for MNIST and CIFAR-10 datasets and show that Bayesian emsembling can improve classification accuracy. For the sake of generality we consider low-rank matrix factorization problem and also show that Bayesian ensembling here can improve prediction quality.
@xsect
In this section we first review the conventional Hamiltonian Monte-Carlo approach and discuss some properties of orthogonal matrices manifold. Then we describe our method called Orthogonal HMC (oHMC) and Orthogonal Stochastic Gradient HMC (oSGHMC) for non-stochastic and stochastic regimes correspondingly.
@xsect
Suppose we want to sample from distribution Ï€(θ)âˆexp(−U(θ)) which we know only up to normalizing constant. For example, we may want to sample from posterior distribution of model parameters given data, i.e. U(θ)=−logp(Data∣θ)−logp(θ). In this situation we can resort to Hamiltonian Monte-Carlo approach that is essentially a Metropolis algorithm with special proposal distribution constructed using Hamiltonian dynamics. Let’s consider the following joint distribution of θ and auxiliary variable r:
The function H(θ,r) is called Hamiltonian. Then for sampling from π(θ) it is enough to sample from the joint distribution π(θ,r) and then discard auxiliary variable r. As joint distribution factorizes
we can sample r∼N(0,M) and then sample θ from π(θ∣r). To do this we simulate the Hamiltonian dynamics:
It can be shown that solution of these differential equations preserves the value of Hamiltonian, i.e. H(θ(t),r(t))=const. Hence we can start from r(0)=r and θ(0) equaling to previous θ, take (θ(t),r(t)) from some t as new proposal and accept/reject it using standard Metropolis formula. Since H(θ(t),r(t))=H(θ(0),r(0)) the new point will be accepted with probability paccept=1.
In practice differential equations (1)-(2) cannot be integrated analytically for arbitrary distributions π(θ). However, there exist a special class of numerical integrators for Hamiltonian equations which are called symplectic. Such methods are known to approximately conserve Hamiltonian of the system and produce quite accurate solutions (Hairer, 2006). The most popular symplectic method for HMC is Leapfrog. One iteration of this method looks as follows:
Since numerical methods introduce some error, we need to calculate Metropolis acceptance probability using old and new Hamiltonian:
If π(θ) is posterior distribution of model parameters θ given Data={xi}Ni=1, then ∇θlogπ(θ) can be written as follows:
When N is big enough the sum becomes hard to compute. The straightforward idea is to compute stochastic gradient over a minibatch instead:
However, this introduces noise in the Hamiltonian equations and forces to change integration method to counteract this. Moreover, exact calculation of Metropolis acceptance probability becomes intractable since we need to sum log-likelihood over all the objects. This forces to use very small step size to make acceptance probability approximately one. Stochastic algorithms can be found in Chen et al. (2014).
Suppose now that we want to sample from distribution π(θ), but θ lies on a Riemannian manifold. This means that we are given a Riemannian tensor G(θ) defining the inner product at every point θ:
This inner product has direct influence on how distances are computed in the θ space and this should be taken into account in efficient sampler in order to make larger steps in parameter space. This was done in Girolami & Calderhead (2011) and in Ma et al. (2015). The main idea is to adapt the Hamiltonian to account for Riemannian structure:
Also, in this case the Leapfrog should be replaced with the Generalized Leapfrog.
It’s important to note that both conventional HMC and Riemannian HMC treat parameters θ and r as unconstrained variables. If this is not true, HMC steps could move parameters off the manifold. Let’s consider an example, when we have some distribution π(e), e∈E, where E is two-dimensional ellipse with axes 2a and 2b. We can parameterize the points on the ellipse at least in two ways:
In the first case we can apply HMC or Riemannian HMC directly since variable t and auxiliary variable are unconstrained. In this case G(t)=a2sin2t+b2cos2t. In the second case we should have (x,y) lying on the ellipse. As auxiliary variable r is, in a sense, the point velocity it cannot have non-zero component orthogonal to the tangent space. Moreover, even if r lies in the tangent space, modifying (x,y) using equation (4) would move the point off the manifold.
For the case of rectangular orthogonal matrices it is hard to find unambiguous unconstrained parameterization. It means that for sampling on this manifold we need to do iterations that 1) always leave θ on the manifold and 2) forces r to lie in the tangent space for the current point θ. Hopefully, this can be done using known properties of orthogonal matrix manifold.
@xsect@xsect
In this subsection we present some useful definitions and concepts about manifolds. More complete discussion of this topic can be found in Absil & Malick (2012).
There is a concept of straight line in Euclidean space. On the manifold this concept is extended using notion of geodesic curve which is defined as curve γ(t) which locally minimizes arc length. Given smooth manifold M there exists tangent space TXM at every point X∈M. Given X∈M,U∈TXM,t∈R we can define exponential mapExp(X,U) as γ(1)=Γ(1,X,U) where Γ(t,x,u) is defined as a curve with the properties:
It is known that Exp(X,tU)=Γ(t,X,U). In Byrne & Girolami (2013) authors use exponential map to define the sampling scheme. However, in many cases exact geodesics are quite hard to compute. Any map R(t,X,U)=R(X,tU) which satisfies equations (6) is called retraction. Retractions are known to be (at least) first-order approximations of exponential map. Usually, retraction is less expensive computationally than corresponding exponential map.
Along with retraction there is also a concept of vector transport. Intuitively, when some point on the manifold moves along a curve its tangent space and all vectors in it are also moving. This process is described via notion of vector transport. Formally, for X,Y∈M it is defined as a function TX→Y:TXM→TYM, which is linear in its argument. A canonical example of such transformation is so-called parallel transport. More information can be found in Sato & Iwai (2015).
@xsect
In this subsection we present some properties of the manifold of the orthogonal matrices that will be useful later. Many of them can be found in Tagare (2011).
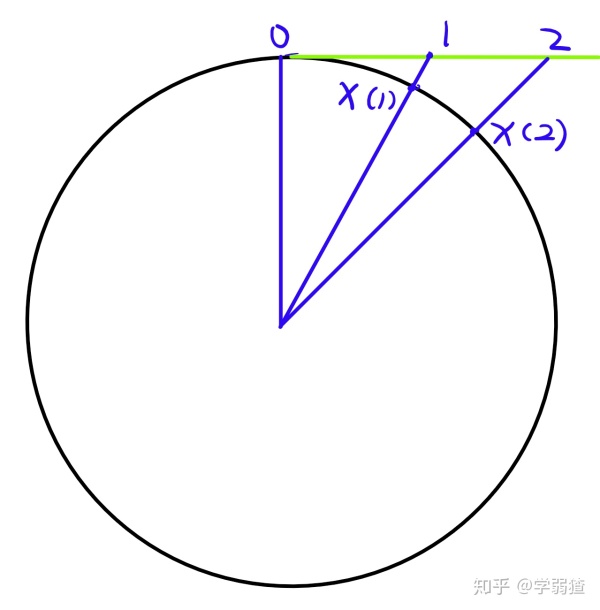
Let Vp(Rn)={X∈Rn×p∣XTX=I} where n≥p denote the Stiefel manifold. There is a convenient representation for the tangent space:
where X⊥ is an orthogonal complement to a set of orthogonal vectors X.
For arbitrary matrix G∈Rn×p it is possible to find projection of this matrix to the tangent space at point X analytically:
Projection is taken with respect to the canonical inner product:
For the Stiefel manifold exponential map and parallel transport for t:(X,U)→(X′,U′) are computed simultaneously as follows (Byrne & Girolami, 2013):
where A=XTU=−UTX,S=UTU.Time complexity of that update is at least two matrix exponentials of size p×p which can be computed in O(p3) time plus time complexity of matrix multiplication which is O(np2) in this case. While in the case of big n and small p computation time for matrix exponentials is almost negligible, it is not the case for the square matrices.
Finally, for Stiefel manifold there are many examples of retractions. Most of them can be found in Absil et al. (2009). One particularly important example is based on Cayley transform.Let AT=−A∈Rn×n,X∈Vp(Rn). Let
Then it is true that:
Thus, if we take A=−(GXT−XGT), then
This retraction is useful because it makes our sampling scheme symplectic which is desirable.
@xsect
Suppose we want to sample from distribution π(θ) of orthogonal matrices. The main idea is to try to adapt the Leapfrog integration (3)-(5) in order to maintain orthogonality constraint for θ and importantly, preserve symplecticity of update. At first, we note that at every iteration we should haveθ∈Vp(Rn) and r∈TθVp(Rn). Secondly, we note that if θ∈Vp(Rn) then Riemannian gradient ^∇θlogπ(θ)∈TθVp(Rn) is definedas a particular vector in the tangent space. Given that the update (3) would keep momentum r in the same tangent space. For updating θ we can use retraction defined in equation (8). So, the update formula (4) transforms to the following one:
Yet, that inevitably changes the tangent space and we lose the property that r∈TθVp(Rn). Nonetheless, we can fix this by making vector transport for r to the new tangent space. An orthogonal matrix X∈Vp(Rn) is an orthonormal system of p vectors on a n-dimensional sphere. The tangent matrix Z∈TXVp(Rn) is a matrix of tangent vectors which show the direction of rotation. Apparently, if the points on the sphere are rotated in some direction then their velocities should be rotated in the same direction too. So, θ and r should be updated at the same time to make sure r∈TθVp(Rn).Finally we get the following update:
Combining these update rules we come to the final scheme:
This scheme is symmetric (can be reversed in time) and symplectic. The full proof of symplecticity is given in supplementary material. The sketch of the proof is as follows: first we need to compute Jacobian J=∂(θn+1,rn+1)∂(θn,rn). This can be done using Woodbury identity several times to rewrite inverse matrices. Having this done, we need to prove that JTAJ=A, where
This can be done directly by multiplying these matrices in the block fashion.
In conclusion, algorithm works like HMC, but uses new scheme for orthogonal matrices. Leapfrog consists of three steps, so if there are several groups of parameters, constrained or unconstrained, we can run each of Leapfrog steps independently on each group of parameters. The pseudocode is shown in Algorithm 1. The inverse matrix in equation (10) can be computed more efficiently using Woodbury identity. This allows us to reduce the time complexity from O(n3) to O(p3). The overall time complexity is O(p3+np2)=O(np2). This suggests that it is best to use our method with tall matrices of small rank. While asymptotically this method has the same time complexity as Geodesic Monte-Carlo, in practice it is faster and more stable which is shown in the experiments.
Stochastic version of oHMC is introduced in the same way as in SGHMC replacing the Leapfrog with our scheme. The pseudocode is shown in Algorithm 2.
@xsect
In this section we give experimental results for oHMC and oSGHMC methods. The first experiments are toy ones intended to show that the proposed method truly works and can sample from the correct distribution.Also we compare our method with Geodesic HMC (Byrne & Girolami, 2013) and show its improved time per iteration and numerical stability along with better effective sample size.Next we apply oSGHMC for Bayesian neural networks where parameters in fully-connected and convolutional layers are given by orthogonal matrices. Finally we experiment with Bayesian models consisting of low-rank matrices parameterized by singular value decomposition. Here we consider both simple low-rank matrix factorization model and neural network with low-rank fully connected layers.
@xsect
Here we consider a mixture of matrix normal distributions as a distribution to sample from. To introduce orthogonal matrices we represent the matrix using QR-decomposition. In other words, the distribution is defined in the following way:
where Q∈Rn×p is orthogonal and R∈Rp×p is upper-triangular.We choose the following parameters: m=16,πi=1m,σ=0.3. We consider two setups: n=p=2 and n=3,p=2 to show that our method works both for square and rectangular matrices. Each mode Mi is a matrix where any element Mijk is 1 or 2. Initial point for sampling is equal to M1.We run oHMC for the nsamples=20000 samples skipping the first nburn=10000 samples.Then we plot every tenth sample on some pair of coordinates. The result is shown in figure 1. Results for square and rectangular matrices are not too different and we show them only for square matrices. To get samples from the true distribution we just sample from the usual matrix normal mixture and then compute QR-decomposition of every sample matrix. As can be seen from the figure, all the modes are covered with sampled points.
We also compare our method with HMC where Q is parameterized as Q=X(XTX)−1/2. This is ambiguous parameterization which can possibly diminish efficiency of the sampler. We compare our method with Geodesic HMC as well. All methods cover the density well enough, however, our method achieves better Effective Sample Size (ESS) which is reported in table 1. Time per iteration of our method is 1.5 times better than of Geodesic HMC and comparable with standard HMC as reported in table 2.
The reason why Geodesic HMC is slower is because of the need to compute matrix exponential twice (7). While time complexity of this operation is asymptotically the same as of matrix inverse, in fact it is more expensive computationally. Another issue with matrix exponential is that it is not quite stable numerically. In this experiment, we could not use larger step size ε for Geodesic HMC because matrix exponential computation failed. Therefore effective sample size for gHMC is less than for oHMC.
@xsect
In this experiment we validate that the stochastic version also works correctly. We take the same model as in the previous experiment, but consider unimodal distribution with m=1,σ=1. We run both oHMC and oSGHMC. For the stochastic method we add artificial noise to the gradients:
where σnoise=0.1. As stochastic method needs more iterations to explore the distribution we take nsamples=50000,nburn=10000. Results are shown in figure 2. As we can see, stochastic method still covers the density well and there is no collapse to the mode.
We also compare SGHMC with oSGHMC in terms of ESS. Our method achieves significantly better ESS which shows that our method explores the distribution faster. Results are given in table 3.
@xsect
In this experiment we demonstrate that oSGHMC can be applied also to neural networks. For this we train orthogonal VGG-16 (Simonyan & Zisserman, 2014) on CIFAR-10 and a small orthogonal multilayer perceptron (MLP) on MNIST. By orthogonal network we mean the network where all weight matrices except for the one from the last layer are replaced with orthogonal ones. For convolutional kernel W∈Ro×i×w×h we reshape it to matrix W∈Ro×iwh and require W to be orthogonal.To make sure the performance of sampling and not optimization is measured, we first find good initial point to start sampler from. To do this we optimize the network using oHMC method, where we switch noise to zero. So in fact the method turns into SGD + momentum-like optimization method over orthogonal matrices.
For MNIST dataset we take an orthogonal MLP with the following architecture: 784-300-300-10, where each number shows the number of units on each layer. The first number is the dimensionality of input. The last layer is the usual (non-orthogonal) fully-connected layer. Such choice is motivated by the fact that since orthogonal matrices preserve norm of the input we need to be able to scale them arbitrarily before the softmax. That is, if we make the last fully-connected layer to be orthogonal, then accuracy significantly reduces which proves our intuition.
We run the optimization for 50 epochs and achieve 97.26% accuracy on the test set. After that we run oSGHMC for nsamples=25000 samples discarding the first nburn=5000. For Bayesian ensemble we take each 1000th sample. Other parameters are: ε=0.00015, batch size=256. The results are given in table 4.
For CIFAR-10 we run optimization of VGG-16 for 200 epochs and get 92.70±0.05% accuracy on the test set. In comparison, for the same VGG-16 but with usual weight matrices we need 300 epochs to get the same quality. We want to note that our network has reduced capacity comparing to usual VGG-16 because of orthogonality constraints. Nevertheless, it can achieve almost the same quality. We take learning rate ε=0.1 and divide it by 10 after 150 epochs. We choose batch size = 128.
After that we run our sampler for nall=24000 samples where we discard first nburn=4000 as burn-in and take the last nsamples=20000. We take each 2000th sample to form an ensemble of 10 networks. Using the ensemble we get 92.85±0.03% test accuracy. Here we take rather small ε=0.00001 to make sure the method does not diverge. We gather all results in table 5. Results for non-orthogonal networks are taken from Izmailov et al. (2018).
@xsect
In this experiment we show how we can use orthogonal parameters to parameterize low-rank matrices.The straightforward way to get a low-rank matrix W∈Rm×n is to make a bottleneck:
This factorization is ambiguous since multiplying W1 by a constant and dividing W2 by the same constant leaves W unchanged. This property makes optimization and sampling harder. On the other hand, there exists unique factorization:
This is well-known singular value decomposition (SVD). This parameterization is non-redundant and can be much more efficient for optimization and sampling.
We apply these two methods to factorize the rating matrix from the MovieLens-100k dataset. We take the rank=10. The preprocessing goes as follows: we center all ratings to lie in [−2,2] where new rating ^r is defined as ^r=r−3. After that we run optimization to find a good starting point for the sampler. Afterwards, we run our sampler for nsamples=5000 with ε=0.0001. We discard the first nburn=1000 samples. We take each 500th sample and get an ensemble of 10 factorizations. The results are collected in table 6.
@xsect
In this experiment we show that low-rank matrix parameterization is applicable for the deep models too.Suppose we want to have linear layers with low-rank weight matrices. We may want to do this because this reduces number of layer parameters and allows to make our network smaller hopefully without quality deterioration.
The dataset of choice here is MNIST which is a dataset of 60000 images of size 28x28. The train part is the first 50000 images and the rest is the test part. We construct a multi-layer perceptron (MLP) using only low-rank layers. The architecture is as follows: we have two hidden layers with size of 784 which is quite big, however for the first layer we upper bound the rank by 100 and for the second by 10. The output layer is full-rank. That is, our architecture can be written as 784-784,100-784,10-10,10. We test our method against the same perceptron but in bottleneck parametrization. We train both networks for 20 epochs. The results on the train/test set are given in table 7. The accuracy is quite reasonable for the network without convolutional layers. Moreover, we can see in figure 3 that in SVD parameterization the training goes much faster.
We also ran sampling from posterior distribution for our network, however, it did not help much, probably because the accuracy is already quite high for MLP.
@xsect
In this paper we have considered a new sampling scheme for the manifold of orthogonal matrices. From the results, it is clear that oHMC improves effective sample size comparing to other methods. We have showed that our method is applicable not only for toy problems, but also for the harder tasks such as Bayesian inference for neural networks.
One of the possible concerns about the proposed method is the irreducibility property. Roughly speaking, this property means that the Markov Chain which the method constructs can travel through the whole parameter space. For the square matrices it is quite obvious that our method is not irreducible since the retraction step X→R(X,Z) preserves detX but real orthogonal matrices have two possible determinants: 1 and −1. This issue can be resolved by working with unitary matrices as they constitute a connected manifold.
For the rectangular matrices the answer is not known for us. However, there is the following intuition: for any rectangular orthogonal matrix X we can find an orthogonal complement such that the resulting square orthogonal matrix will have the determinant of desired sign. Therefore, any two rectangular can be connected using our retraction. Although this is not a formal proof, it gives us some hints about the answer to the question about the irreducibility of the method.
@xsect
In the paper we have presented a new HMC sampler for Bayesian models with orthogonal matrices. Experimental results have shown that new sampling scheme can be much more sample-efficient comparing to conventional HMC with ambiguous parameterization for orthogonal matrices. Using proposed method it becomes possible to make Bayesian ensembling for orthogonal deep neural networks.
Orthogonal matrices can also be used in parameterizations for low-rank matrices, positive-definite matrices and others. It is known that a set of low-rank matrices forms a Riemannian manifold (Vandereycken, 2013). It should be noted that the proposed oHMC and oSGHMC methods do not fully account for this manifold since here matrices U and V in SVD parameterization are allowed to change independently. We consider finding the optimal Riemannian HMC sampler for low-rank matrices for future work. In the same way it is interesting to generalize our sampling scheme for tensors in tensor train format (Steinlechner, 2015). This family for tensor representation becomes more and more popular and, in particular, can be used within neural networks (Novikov et al., 2015).
References
For questions about Stiefel-manifolds, the set of all orthonormal k-frames in R^n.
2
3answers
How to solve the system of matrix equations $XX^TA = A$, $X^TX = I$?
Given tall matrix $A in mathbb R^{n times k}$ (where $n gg k$), is there a way to solve the following system of matrix equations in $X in mathbb R^{n times k}$?$$begin{aligned} X X^T A &=...
2
1answer
Maximize trace over Stiefel manifold
This question is the same as the question in this post. The OP of that post changed what they were asking and reduced it to a special case, so I’m asking the question in full generality here. Given ...
1
1answer
References for injectivity/surjectivity $pi_i(U(n-k)) rightarrow pi_i(U(n))$
I have read (Theorem 29.3, line 7, pg 144) the following statement: $pi_i(U(n-k)) rightarrow pi_i(U(n))$ is surjective for $i le 2(n-k)+1$ and injective for $i le 2(n-k)$. and ...
CL.
2,68433 gold badges99 silver badges2626 bronze badges
2
1answer
Smallest trace of a matrix product where one is given and the other is orthogonal
What are the optimal solution and optimal value for the following semidefinite program$$ min_{ V } { mbox{tr} (VSigma) : VV^T=I }$$where $Sigma$ is a given positive semidefinite matrix, and ...
3
1answer
How to optimize objective in the Grassmann manifold?
For Stiefel manifold, it contains all the orthogonal column matrices$$St(d,M) = {X in R^{M times d} | X^TX = I}$$For Grassmann manifold, it is $$Gr(d,M) = {col(X), X in R^{M times d}}$$...
1
0answers
'Jacobian' of QR decomposition of a rectangular matrix
I want to calculate the volume of real Stiefel manifold $V_{k}(mathbb{R}^N)$ .$$ V_{k} (mathbb{R}^N) = { H in M(N, k, mathbb{R})| H^{T}H = I_{k} }$$((^T) denotes transposed matrix. $M(N, k, ...
0
0answers
Closed geodesics on the Stiefel manifold
I would like a reference on the following property of Stiefel manifolds $St(p,n)$: A closed geodesic $c:[a,b]rightarrow St(p,n)$ is a local minimum of the curve length function$$L(gamma)=int_a^b|...
1
0answers
General Stiefel-Whitney classes and Stiefel manifolds
Here are some statements that I wish to understand more deeply, whose truth value I want to check, and to determine under which criteria they are valid. Consider the Stiefel manifold $V_k(R^n)$ of ...
wonderich
2,38833 gold badges1414 silver badges3131 bronze badges
3
1answer
Spheres and orthogonal matrices as spaces of solutions to matrix equations
For $a in mathbb{R}^n$, the solutions to $$1=sum_{i=1}^na_i^2$$form an $(n-1)$-sphere in $mathbb{R}^n$. Meanwhile, for $A in mbox{GL}_k (mathbb{R})$, the solutions to$$1=AA^T$$are the ...
2
1answer
BO(-) example in Weiss Calculus
I'nm reading Orthogonal Calculus by Michael Weiss, and trying to understand example 2.7, concerning the derivatives of the functor $BO$, which sends a (finite dimensional) inner product space to the ...
3
1answer
Minimize $ mbox{tr} ( X^T A X ) + lambda mbox{tr} ( X^T B ) $ subject to $ X^T X = I $ - Linear Matrix Function with Norm Equality Constraint
We have the following optimization problem in tall matrix $X inmathbb R^{n times k}$$$begin{array}{ll} text{minimize} & mbox{tr}(X^T A X) + lambda ,mbox{tr}(X^T B) text{subject to} &...
1
0answers
From what manifold do we need to start to build an eversion for $any$-tridimensional object?
If a sphere eversion is possible using a half-way model how model is used for $cylinder$ eversion ?I need to make some premises to be able to frame the true nature of the problem In sphere eversion ...
1
0answers
Self maps on Stiefel Manifolds
Let $m_l: S^{n-1} stackrel{z mapsto z^l}{to} S^{n-1}$ be a map of spheres. Then it induces a map on $( S^{n-1})^k$, given by product. Note the Stiefel manifold $V_k(mathbb{R^{n}})$, the ...
3
0answers
Neighbourhood of a point in Stiefel manifold
In Hatcher above, he claims that 'This determines orthonormal bases for the $(k-m)$ planes orthogonal to all nearby $m$-frames', I feel confused about this claim (even after checking the preceding ...
4
1answer
Nearest Semi Orthonormal Matrix Using the Entry Wise $ {ell}_{1} $ Norm
Given an $m times n$ matrix $M$ ($m geq n$), the nearest semi-orthonormal matrix problem in $m times n$ matrix $R$ is$$begin{array}{ll} text{minimize} & | M - R |_F text{subject to} &...
153050per page
Acknowledgements
This paper presents research results of the Belgian Network DYSCO (Dynamical Systems, Control, and Optimization), funded by the Interuniversity Attraction Poles Programme, initiated by the Belgian State, Science Policy Office. The scientific responsibility rests with its authors. This work was supported in part by “Communauté française de Belgique - Actions de Recherche Concertées†and by the Australian Research Council through discovery grant DP0987411 “State Observers for Control Systems with Symmetryâ€.
This work benefited from fruitful interaction with a fairly large number of colleagues, most of whom appear in the bibliography. The first author would like to thank especially Alan Edelman, Uwe Helmke, and Knut Hüper for helping him into this elegant area of research, and Kyle Gallivan and Paul Van Dooren for a valued and productive collaboration.
All manifolds share same API. Some manifols may have several implementations of retraction operation, every implementation has a corresponding class.
Comments are closed.
|
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
- Home
- New Page
- About Me
- Blog
- Amharic Fiction Books Pdf
- Kanye West Drum Kit
- Kingdom Come Playing With The Devil Bug
- Sniper Ghost Warrior Pc Game Free Download
- Eli Kasim Mp3 Download
- Morrowind Merge Leveled Lists
- Partypoker Get Started Mission
- Windows Unable To Complete The Format
- Max Andres Madigo La Hora
- Torchlight 2 Best Build
- Halo Wars 1 Mods
- Diablo 2 Lord Of Destruction Download
- Yu Gi Oh Tag Force Special Update
- Crysis 3 Save File
- Download After Effect Portable 32 Bit Cs5
- Sarhad Paar Movie Free Download
- The Beatles Stereo Box Set Torrent
- Tron Legacy Pc Game Torrent
- Contact
- Fallout New Vegas Save

 RSS Feed
RSS Feed
